8 Week 7- Population Structure & Sea Cucumbers
We will be using data from a paper by Xuereb et al. (2018) paper on P. californicus titled “Asymmetric oceanographic processes mediate connectivity and population genetic structure, as revealed by RADseq, in a highly dispersive marine invertebrate (Parastichopus californicus)”.
SNP data from this study will be used today in order to learn how to identify population structure in sea cucumber populations and make structure plots using tess3r and pophelper. This data is SNP data from 15 individuals per site across 7 collection sites:
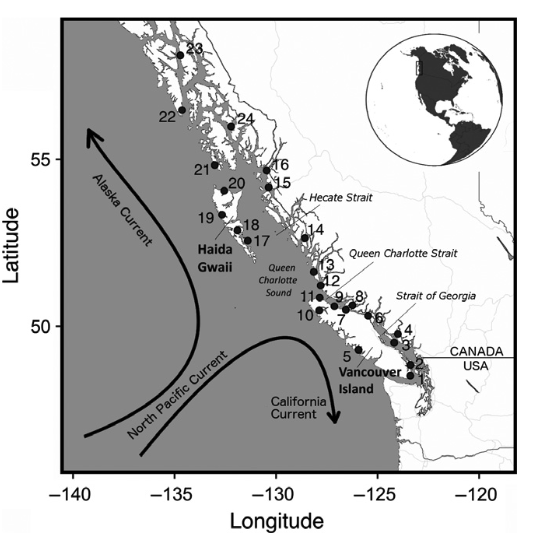
Map of sampling locations in coastal British Columbia (1–20) and southeastern Alaska (21–24).
8.1 Main Objectives:
- Practice moving from terminal to R when working with big data
- Interpret structure plots
- Create structure plots of genomic data in R
8.2 Warmup: Joining command line and R
So, far, we’ve mostly been working with datasets that are available to download in R; but a common practice in genomics is to work with large datasets (like sequencing reads) on a cluster and once the data is processed into a smaller format, working with downstream files in R.
Here is a brief overview of our goals for the warmup:
- Create a new folder in your personal directory on farm called
week7 - Download genomic data into this folder using
wget https://raw.githubusercontent.com/mlarmstrong/IntroGenomics_Data/main/week7.zip - Unzip the data folder and vcf
- Create an RStudio OnDemand Session
- Make a script called
week7and save it in yourweek7directory - Install and load the following R packages:
ggplot2vcfRdevtoolsremotesgridExtradplyr
- Use
devtoolsandremotesto install the following packages from github:
pophelpertess3r
A more thorough walk through of the warm up can be followed below.
8.2.1 Steps 1 through 3: Getting the data onto the cluster
We first need to download the data in terminal. Start an OnDemand session. Navigate to your directory and do the following command to get the vcf file we need onto your farm desktop. We will download a zipped version of this data to make the download easier, but note that you can almost always download published genomic data from online repositories like dryad or NCBI. This data set is available on dryad here: https://datadryad.org/dataset/doi:10.5061/dryad.db6177b
cd /group/rbaygrp/eve198-genomics/yourdirectory
wget https://raw.githubusercontent.com/mlarmstrong/IntroGenomics_Data/main/week7.zipUnzip the week7 directory with unzip week7.zip and navigate inside with cd week7.
There should be three files:
- The README.txt gives you information on the dataset you just downloaded.
- The .csv file in this folder is our metadata, with coordinates for each of our sample sites along with sample names. This will be useful for understanding population structure between populations and across space!
- The vcf file contains SNPs, similar to one we produced with our variant calling
pipeline a few weeks ago! If you file has the extensions .zip, that means that
even though your folder is unzipped, this file is still compressed. You can run
unzip filtered_2719neutral_snps.recode.vcf.zipto unzip it!
8.2.2 Steps 4 through 7: Getting set up in R
Now we will move to R! Start by creating an OnDemand R Studio session.
Go to file>new to create a new R script. Name it week7.R and save it within
the week7 folder you created in your personal directory above.
We will need quite a few packages today since genomic datasets require special tools!
We will be using tess3r to identify population structure and then
pophelper to visualize that structure in a more organized way. We also
need to be able to read in a vcf file, so vcfR is a useful package for
Most packages can be installed using install.packages() and then loaded using
library(). Let’s do this for ggplot2, vcfR, devtools and remotes.
devtools and remotes are R packages that allow us to install even more
packages than can be installed using base R. More on that below!
install.packages(c("vcfR", "ggplot2", "devtools", "remotes", "gridExtra", "dplyr")) #only run this line once, it might take a few minutes
library(vcfR)
library(ggplot2)
library(devtools)
library(gridExtra)
library(dplyr)You can need to use devtools to install tess3r, but then it can be loaded using
library(), just like other packages.
devtools::install_github("bcm-uga/TESS3_encho_sen") #only run once, you can press 'enter' to skip updates
library(tess3r)Similarly to devtools, remotes is a package you can load in R and use to install
other packages that are available on github, like you pophelper
remotes::install_github('royfrancis/pophelper') #only run once, this might take a few minutes!
library(pophelper)8.3 Setting your Working Directory in R
We downloaded data and created a script in the folder called week7, and we want
to make sure that R recognizes this as our working directory. You can think of
this process as similar to when you cd to change directories on the command line.
To set your working directory you can do the following:
setwd("/group/rbaygrp/eve198-genomics/yourpersonalfolder/week7")
A few pro tips:
- You can use tab complete to fill in your path to your folder!
- You can check your current directory with the command
getwd()
8.4 Reading in vcfs
Now let’s read in our data! This will take a second and you will get information on the file in your console as it is read in. The number of variants that are processed should be 2,719. Then you can extract the genotypes with the second line of code.
vcf_cukes<-read.vcfR("filtered_2719neutral_snps.recode.vcf")
genotypes_raw<-extract.gt(vcf_cukes)If you do head(genotypes_raw) you can see that the format is 0/0 or 1/1
for homozygotes and 0/1 for heterozygotes. We actually want to convert
this to instead be 0 for homozyous reference, 1 for heterozygotes and 2
for homozyogous alternate. To do this, you can run the following code:
## Replace genotype codes with numeric values
genotypes_cleaned <- as.matrix(apply(genotypes_raw, c(1, 2), function(x) {
if (x %in% c("0/0", "0|0")) return(0) # Homozygous reference → 0
if (x %in% c("0/1", "1/0", "0|1", "1|0")) return(1) # Heterozygous → 1
if (x %in% c("1/1", "1|1")) return(2) # Homozygous alternate → 2
return(NA) # Handle missing or other cases
}))Check to see that it worked by doing View(genotypes_cleaned) or
head(genotypes_cleaned). The genotypes should be coded as 0, 1 or 2.
Finally let’s check the dimensions of our genotype data. The number of
individuals should be on the left and the genotypes should be on the
right.
dim(genotypes_cleaned) #indiv should be on left, genotypes on rightIt looks like it is actually flipped, so let’s fix that! The command
t() will help us transpose our data
genotypes<- t(genotypes_cleaned)
dim(genotypes) #now it is right!8.5 Reading in csvs
Now let’s read in the coordinate data for our sample sites! We will also
make an object “coordinates” to hold our lat/long data for the tess3
object later. Our sites.data should a length of 717 to match our
genotype data.
sites.data<-read.csv('strata.cucumbers.717ind.csv', header=TRUE, sep=",")
coordinates<-as.matrix(cbind(sites.data$LAT, sites.data$LONG))
dim(sites.data)8.6 Mini Lecture: Interpreting Structure Plots
Let’s spend a bit more time getting oriented to the sea cucumber paper and understanding more generally what information is represented with structure plots!
You can follow along with the slides here
8.7 population structure with tess3r
You can learn more about this package on their tutorial page (https://bcm-uga.github.io/TESS3_encho_sen/articles/main-vignette.html), but in short tess3 is a function that computes population structure by estimating ancestry proportions and ancestral allele frequencies. In the code below: - X argument refers to the genotype matrix - coord argument the coord argument corresponds to the geographic coordinates - K is the number of clusters or ancestral population - reps is the number of replications for cross validation that we are calling the ancestry correctly for each sample
Normally we would want multiple reps to ensure things are being calculated correctly, but for class we will only do 2.
tess3.cukes <- tess3(X=genotypes, coord =as.matrix(cbind(sites.data$LAT, sites.data$LONG)), K=1:8,
ploidy=2, lambda=0, keep="best",rep=2)This will take a second since it is real genomic data! We started with 8 clusters.
Next to make a structure plot, we need to first call the data and assign how many K “clusters” we want. For example, if we want to assign 8 clusters we would run the following:
# retrieve tess3 Q matrix for K = 8 clusters
cukes.matrix <- qmatrix(tess3.cukes, K = 8)#now the barplot
barplot(cukes.matrix, border = NA, space = 0,
xlab = "Individuals", ylab = "Ancestry proportions",
main = "Ancestry matrix") -> bp
axis(1, at = 1:nrow(cukes.matrix), labels = sites.data$Sample,las = 3, cex.axis = .4) Yay our first structure plot! But it isn’t ordered by site (yet!), so it’s hard to see if there are differences across sites.
8.8 How do we know which K to pick?
In order to get an accurate estimate of the “best” K for our data we first need to run a larger range of K, here 1:8, and then look at our data and see how the cross-validation score compares to the number of predicted ancestral populations. We can do that with the code below:
The plot function generates a plot for root mean-squared errors computed on a subset of loci used for cross-validation:
plot(tess3.cukes, pch = 19, col = "orange",
xlab = "Number of ancestral populations",
ylab = "Cross-validation score") Class Exercise 1
Where the cross-validation score drops is which K is likely the best fit. Which K do you think is the best fit for our data? Let’s plot a couple different structure plots (K=2, K=4 and K=6) for data visualization and determine the best K. You can modify the code above by changing the K value and rerunning the plots.
Now that we have determined the best K for our dataset, let’s visualize things!
8.9 Visualizing population structure with pophelper!
Pophelper is a really useful tool for visualizing population structure graphs. You can view the tutorial here for more information: https://www.royfrancis.com/pophelper/articles/index.html#plotting-1. We can do a lot of customization in pophelper, and this figure shows what our final result will look like!
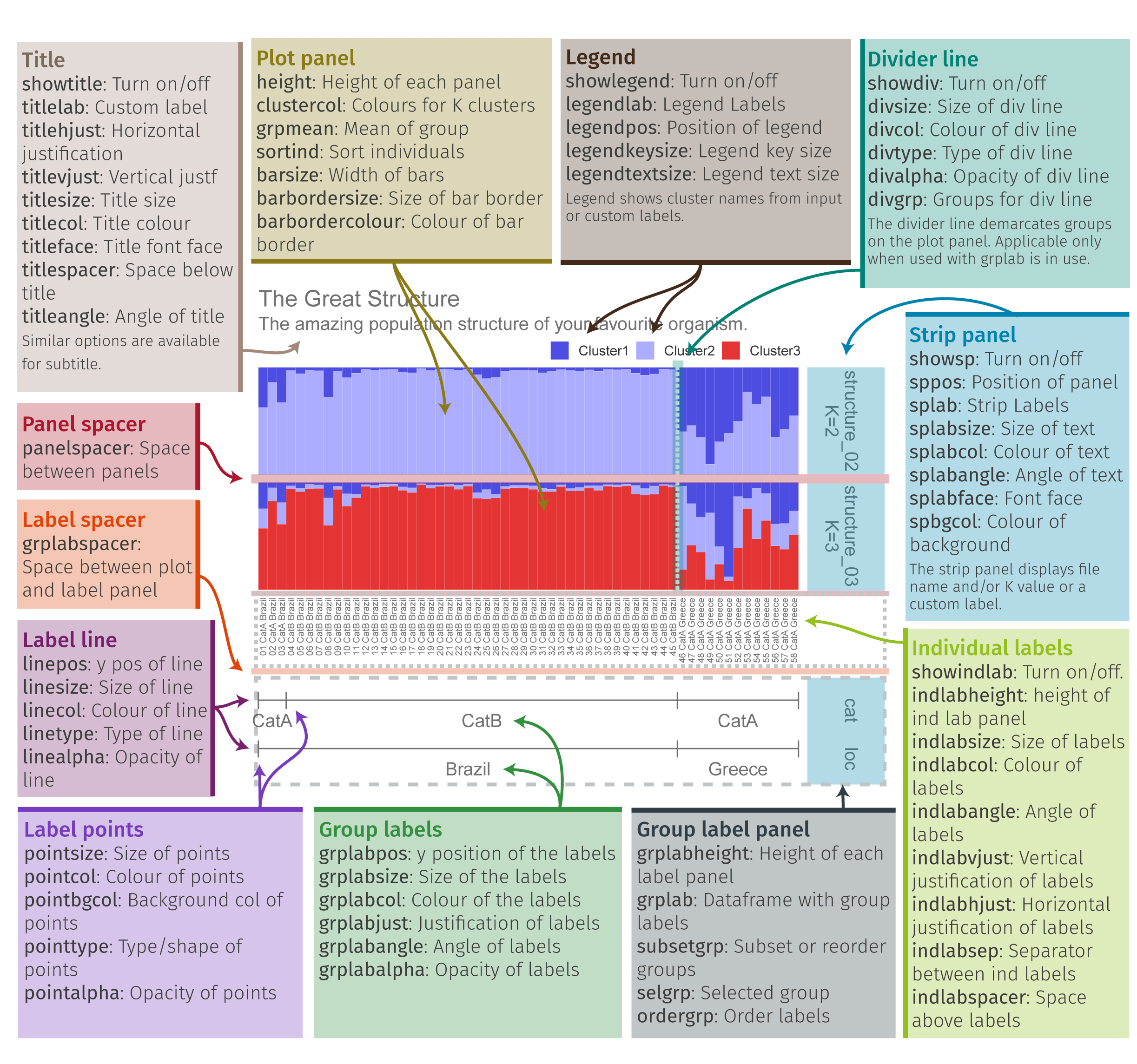
Pophelper Overview
- First we need to convert our tess3 object to a qlist for pophelper
qlist.cukes <- readQTess3(t3list=tess3.cukes)- Then we need to specify the order that we want our sites to be in and the different labels for our structure plot. The first is our population order. We want it to match the map figure and the table in the paper, so let’s order from South to North. To get the same of the sites we can do the following:
unique(sites.data$SITE)Our sites are “OGD” “SGI” “LAS” “JER” “TOF” “CRA” “RBY” “SHE” “MAL” “QUA” “HOP” “TBL” “CAL” “TOL” “PRI” “LEG” “JUA” “SEL” “REN” “MAZ” “AK1” “AK2” “AK3” “AK4”.
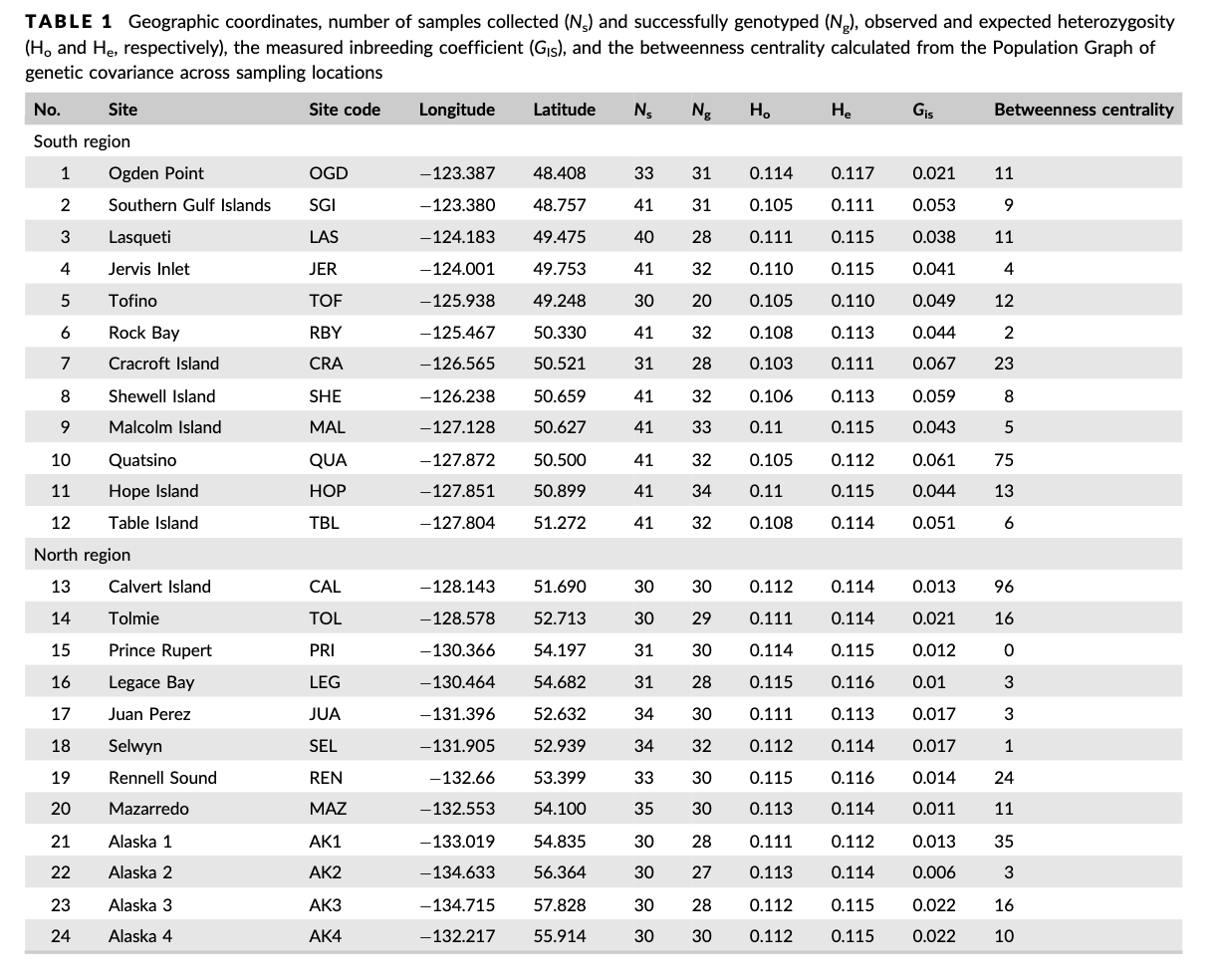
Table 1 from Xuereb et al., 2018
Table 1 from the paper shows the sites south to north. Luckily the dataset is already sorted appropriately, so now we can just put this into a new object!
ordered.sites=c("OGD", "SGI", "LAS" ,"JER" ,"TOF" ,"CRA" ,"RBY", "SHE", "MAL", "QUA", "HOP", "TBL", "CAL", "TOL", "PRI", "LEG", "JUA", "SEL", "REN", "MAZ", "AK1", "AK2" ,"AK3", "AK4")We also want to label our two structure plots with what K equals, so add that with the object “labels”
labels <- c("K=2", "K=3")- Now that we have our labels we can pull the data that we want from our different files
slist.cukes <- alignK(qlist.cukes[2:3],type="auto")
labset<-data.frame(sites.data)- Now we can plot our data!
#plot samples separated by site
p<-plotQ(slist.cukes,imgoutput="join",returnplot=TRUE,exportplot=FALSE,basesize=11,
splabsize=7,height=7,
grplab=data.frame(SITE=labset$SITE),subsetgrp=ordered.sites,
grplabsize=3,linesize=1,pointsize=3,splab=labels,grplabangle=0,
grplabheight = 5)grid.arrange(p$plot[[1]])8.10 Manipulating our metadata with dplyr
The site names may mean something if you are familiar with this area or
have read the paper, but how can we make our plot easier to understand?
One way could be to add what countries these samples were taken in so
you can get a sense of sampling area. To work with this dataset, we’ll use a
really useful package for data manipulation that we haven’t used yet, called dplyr().
The newest thing that you will see with dplyr is %>%. This is called a
pipe, and connects lines of code together.
To add a new column we will need to define what the new column will be named and then provide information to fill that column
library(dplyr)
sites.data<-sites.data %>%
mutate(Country=case_when(
SITE %in% c("OGD", "SGI", "LAS" ,"JER" ,"TOF" ,"CRA" ,"RBY", "SHE", "MAL", "QUA", "HOP", "TBL", "CAL", "TOL", "PRI", "LEG", "JUA", "SEL", "REN", "MAZ") ~"British Columbia, Canada",
SITE %in% c("AK1", "AK2" ,"AK3", "AK4") ~ "Alaska, USA"))View sites.data before moving on to ensure you got the column!
Class Exercise 2
Now we can rerun some of the code we have above to remake our structure plot with “Site” and “Country” labels.
8.11 Group Work Activity- Adding even more context to our plot
The structure plot is now more organized with samples from south to north, but what if we want clear labels for how those sites split? Remember that Table 1 grouped sites by region, with Ogden Point to Table Island as the “South Region” and Calvert Island to Alaska site 4 as the “North Region”.

Table 1 from Xuereb et al., 2018
- Add a column for “Region” in sites.data dataset.
- Make a new structure plot with a label for
SITEandRegion. - Modify the color scheme however you’d like (but make it distinct from the preset
colors, please!) View the pophelper page for more information on how to do so.
Think of how you can use color to convey a message with your plot.
You can check out a few color schemes from
pophelper()here
Submit your code and a photo of your structure plot to canvas.
8.12 Key Points
- Moving from terminal to R is important when working with genomic data
- Structure plots can be made using tess3r and are useful for visualizing population structure within your dataset
- Pophelper is a useful tool for customizing these structure plots
Class Exercise Solutions
Class Exercises: Solutions
Exercise 1
It looks like there might just be two clusters, since the cross-validation score drops between 1 and 2 ancestral populations. Let’s make a matrix using qmatrix for K=2 and visualize a structure plot with that. The qmatrix is a matrix of our ancestry coefficients, aka the matrix of relatedness between our samples! Our qmatrix will be called
cukes.matrix
# retrieve tess3 Q matrix for K = 2 clusters
cukes.matrix <- qmatrix(tess3.cukes, K = 2)#now the barplot
barplot(cukes.matrix, border = NA, space = 0,
xlab = "Individuals", ylab = "Ancestry proportions",
main = "Ancestry matrix") -> bp
axis(1, at = 1:nrow(cukes.matrix), labels = sites.data$Sample,las = 3, cex.axis = .4) Exercise 2
structure plot with site and country labels
labset<-data.frame(sites.data)
verifyGrplab(grplab=labset[,c("SITE", "Country")]) # to make sure it is there
p2<-plotQ(slist.cukes,imgoutput="join",returnplot=TRUE,exportplot=FALSE,basesize=11,
splabsize=7,height=7,
grplab=labset[,c("SITE", "Country")],subsetgrp=ordered.sites,
grplabsize=3,linesize=1,pointsize=3,splab=labels,grplabangle=0,
grplabheight = 5)grid.arrange(p2$plot[[1]])